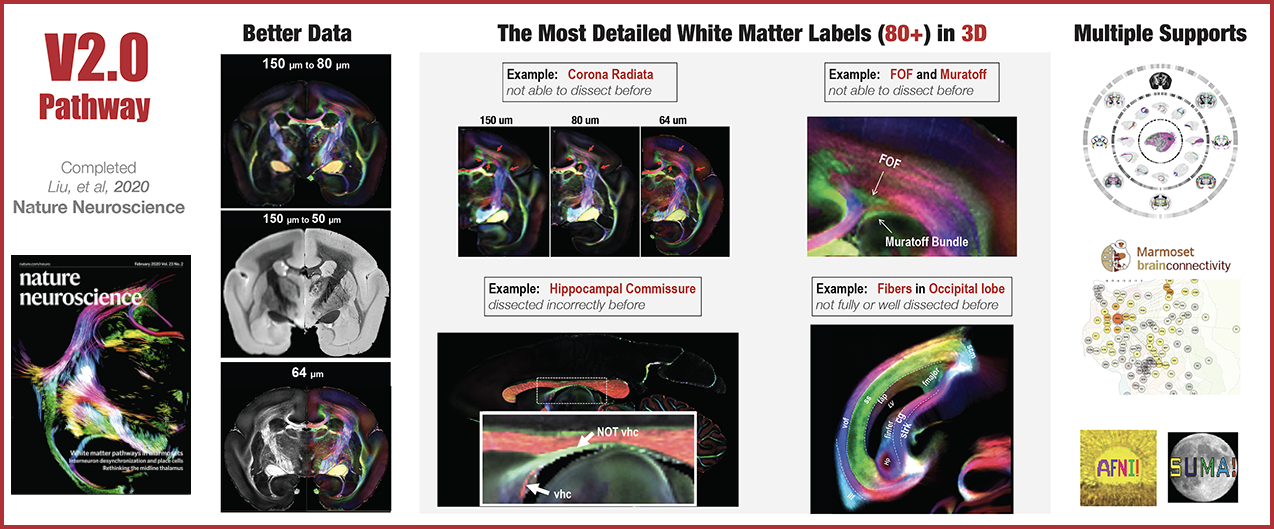
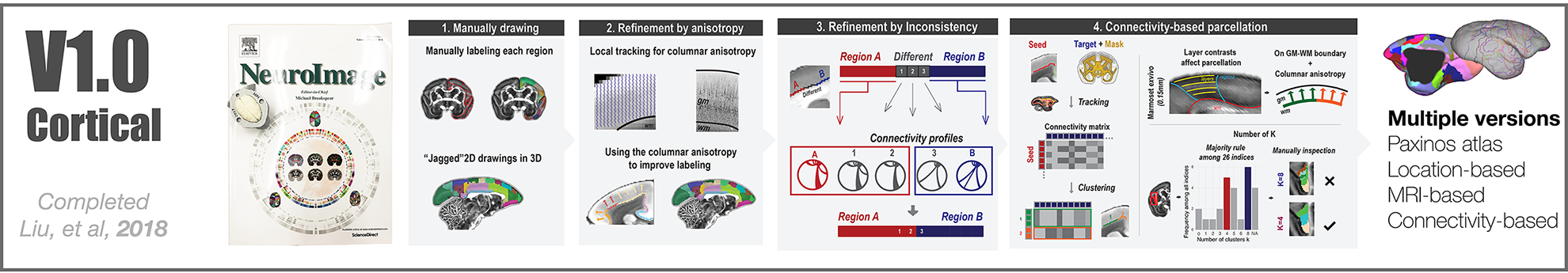
The Version-6 Focused on the Opposing Gradient Axis
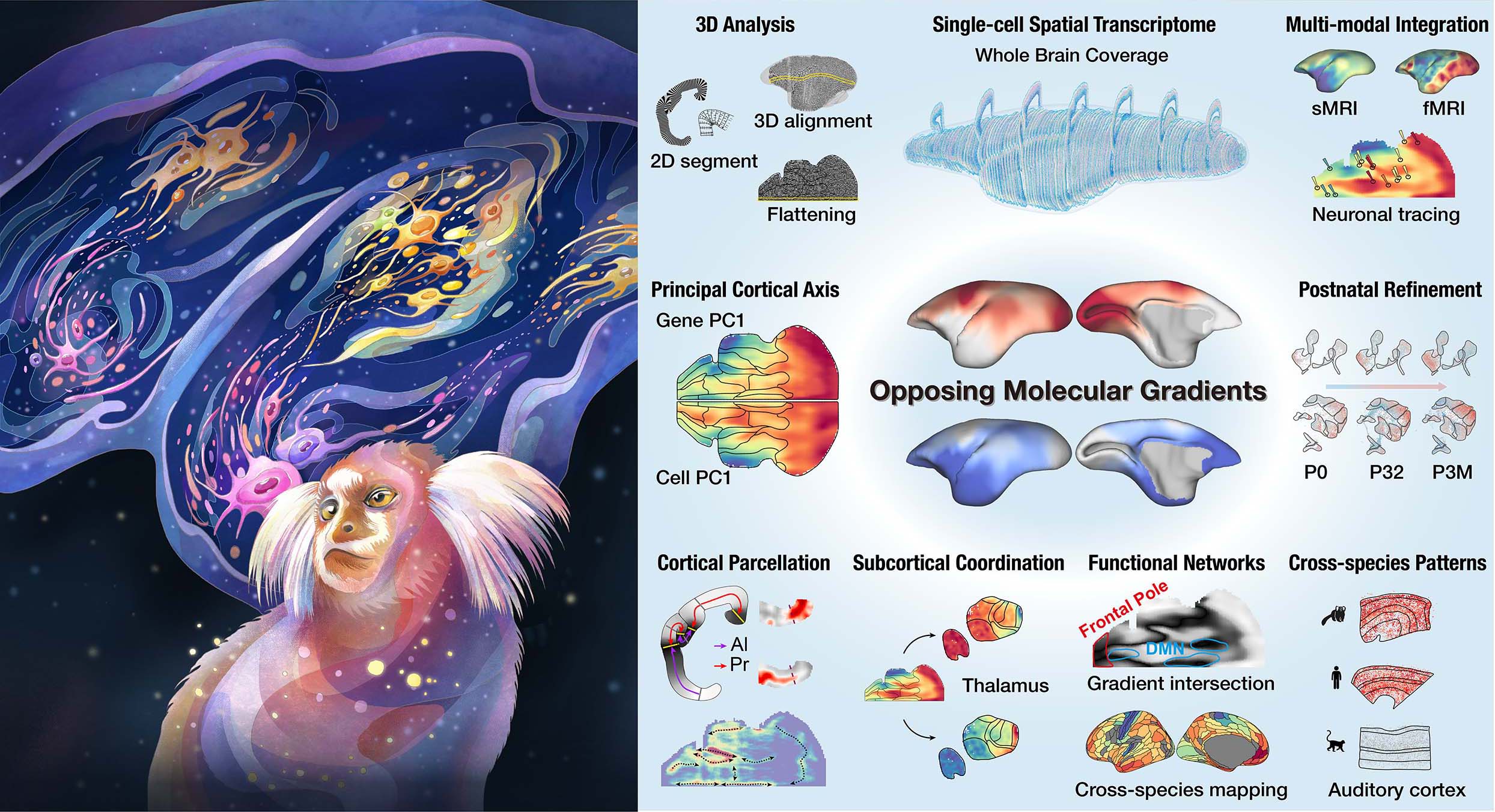
Abstract The principles organizing cellular diversity and connectivity in primate brains remain elusive. By integrating spatial transcriptomics, magnetic resonance imaging, and retrograde labeling in marmosets, we identified two opposing molecular gradients that undergo postnatal refinement, emanating from allocortices and primary sensory cortices, respectively. These gradients reconcile conflicting hypotheses on cortical expansion and characterize distinct cortical areas. Cortical gradients align with thalamic gene expression and thalamocortical projection patterns. At gradient intersections, the default-mode network and frontal pole exhibited similar molecular features in humans and marmosets, despite species differences in functional connectivity. Comparative analysis of gradient-related genes showed that marmoset and human auditory cortices are highly similar but differ from macaques, potentially reflecting complex vocalization. Together, these opposing gradients represent a fundamental organizing principle of primate cortex.